Wei Xie’s group and Bing Zhu’s group published research in Nature Genetics on the mechanisms underlying robust-yet-flexible DNA methylation inheritance
A joint team led by Dr. Wei Xie from School of Life Sciences of at Tsinghua University and Dr. Bing Zhu from Institute of Biophysics at Chinese Academy of Sciences, has revealed how the DNA methylome is stably inherited despite constant epigenetic drift. Their study, titled “Imprecise DNMT1 activity coupled with neighbor-guided correction enables robust yet flexible epigenetic inheritance”, is published in Nature Genetics on July 20, 2020. The research team show that epigenetic drift during cell proliferation in part is due to imprecise DNMT1 activity including both weak de novo activity and imperfect maintenance activity, which leads to spontaneous “epimutations”. However, those epimutations tend to be corrected over time during cell expansion. Interestingly, the correction is likely also executed by the same imprecise DNMT1 activity. Furthermore, such imprecise DNMT1 activity could render the propagation of the methylome from different starting methylation levels, thus enabling the robust-yet-flexible epigenetic inheritance. One key mechanism underlying such “robust-yet-flexible epigenetic inheritance” is a “Tuner-Stabilizer” system based on the neighbor methylation sensitive activity of DNMT1. The researchers believe that similar mechanisms may also underlie the inheritance of other epigenetic marks.
During cell division, the transmission of the genome is accompanied by the propagation of the epigenome, a process that is essential for maintaining cellular memories. However, epigenetic inheritance differs from genetic inheritance in two major aspects. First, the inheritance of the epigenome across cell division is imprecise yet robust. Second, the epigenome inheritance is flexible as it can accommodate and then maintain epigenomes with different starting states. These characteristics are fundamental properties of the epigenome that allows transition and subsequent stabilization of cell fates. However, how the epigenome achieves these properties remains unclear. By generating a “DNMT1-only” cell line, the researchers discovered that clonal population of DNMT1-only cells again produces a heterogeneous methylome, which is stably propagated upon cell expansion and differentiation. Mechanistically, DNMT1 has imprecise maintenance activity and likely possesses weak de novo activity, leading to spontaneous “epimutations”. These epimutations, however, tend to be corrected over time in a neighbor DNA methylation-guided manner. Further analyses support a model that such neighbor-guided correction is enabled by neighbor-guided de novo activity (“tuner”) and maintenance activity (“stabilizer”) of DNMT1. Finally, the researchers generated base-resolution map of de novo and maintenance activities of DNMT1, which revealed that H3K9me2/3 marked regions show enhanced de novo activity, while CpG islands have both poor maintenance and de novo activities. In sum, these data suggest the imprecise epigenetic machinery coupled by neighbor-guided correction may be a fundamental mechanism underlying robust-but-flexible epigenetic inheritance and shed light on how DNA methylation states are maintained at distinct regulatory elements.
Dr. Wei Xie from School of Life Science at Tsinghua University and Dr. Bing Zhu from Institute of Biophysics at Chinese Academy of Sciences are the co-corresponding authors of this work. Qiujun Wang (Postdoc fellow from CLS program at Tsinghua university), Guang Yu (Ph.D. student from School of Life Sciences at Tsinghua University), and Xuan Ming (Institute of Biophysics at Chinese Academy of Sciences) are the co-first authors of this work. Prof. Hehuang Xie and Xiguang Xu from Epigenomics and Computational Biology Lab, Biocomplexity Institute of Virginia Tech also makes important contributions to this study. This work is supported by the National Natural Science Foundation of China, the National Key R&D Program of China, Beijing Municipal Science & Technology Commission, THU-PKU Center for Life Sciences and Chinese Academy of Sciences. Prof. Wei Xie is also an HHMI International Research Scholar.
https://www.nature.com/articles/s41588-020-0661-y
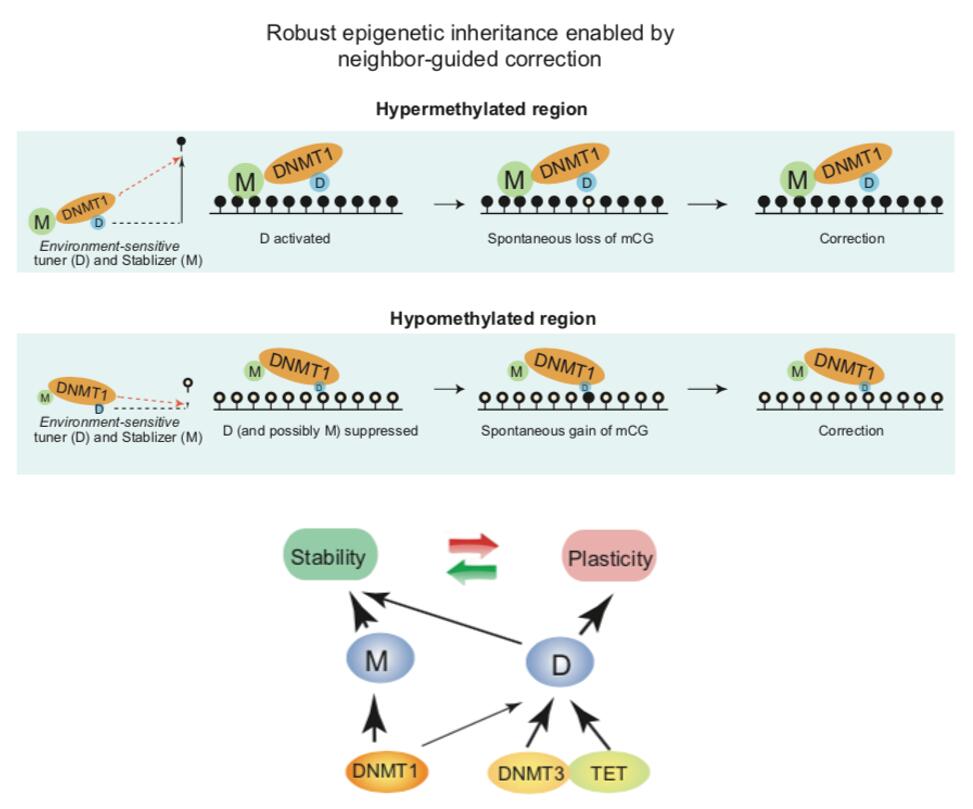
Robust-yet-flexible inheritance of DNA methylation through neighbor-guided correction, enabled by weak de novo activity and imperfect maintenance activity of DNMT1



