By investigating the dynamics of chromatin states during human embryo development, a team led by two groups in Tsinghua University (Wei Xie’s group from CLS and Jie Na’s group from School of Medicine) in close collaboration with Yingpu Sun’s group from the First Affiliated Hospital of Zhengzhou University, has revealed the dynamic chromatin reprogramming process during human preimplantation embryo development. Their findings, published in Nature on May 3rd, 2018, not only advanced the understanding of chromatin reprogramming in human early embryo development, but also provided molecular framework for future studies on Assisted Reproductive Technology (ART) and related human diseases.
Human life starts from a fertilized egg, which undergoes a series of drastic chromatin reprogramming during early embryonic development. In recent years, studies using the mouse model have shown that during early embryonic development, chromatin accessibility, the high order chromatin structure as well as the chromatin modifications undergo tremendous changes on both parental alleles. These changes act in concert with the genome activation and result in a new totipotent embryo, allowing subsequent embryonic development and lineage specification. Previous reports have found that the regulatory elements are usually located in open chromatin. These regulatory elements together with cell type specific transcription factors guide cell fate determination. Analysis of chromatin accessibility enables the identification of regulatory elements and key transcription factors that bind to these elements. However, due to the extremely limited research materials that are available for studying human early development, little is known about the chromatin state and its dynamics in human early embryos. To overcome this hurdle, Wei Xie’s group developed an optimized ATAC-seq protocol that allows chromatin accessibility analysis using as few as 20 cells. By closely collaborating with Yingpu Sun’s group from Center for Reproductive Medicine, the First Affiliated Hospital of Zhengzhou University, the landscape of chromatin accessibility during human early embryonic development was revealed. By working with Jie Na’s group and using mouse embryos, the joint team uncovered both divergent and conservative mechanisms that govern chromatin reprogramming during human and mouse early development.
Through the study of chromatin accessibility in human early embryos, the researchers first identified putative transcription factors that may function during human preimplantation development. By comparing with previous results in mice from Wei Xie’s group, both conserved and species-specific transcription factors were identified. Surprisingly, widespread open chromatin was also found prior to zygotic genome activation (ZGA) in humans (1-4 cell) even the transcription activities are minimal. Many such open chromatin regions reside at CpG rich promoters, and are correlated with genes activated at the 8-cell stage. Unexpectedly, many open chromatin regions also fall into distal regions and are enriched for transcription factor binding sites. Intriguingly, a large fraction of these sites become inaccessible upon the onset of zygotic genome activation. A close examination revealed that these regions also overlap with DNA hypomethylated domains in human oocytes. In mouse, DNA hypomethylated regions in oocytes also preferentially enrich for open chromatin and non-canonical H3K4me3, a unique form of histone modifications that is linked to genome silencing at this developmental stage. The researchers proposed that such pre-ZGA specific open chromatin may serve as “chromatin harbor” for docking or sequestering transcription factors. Closing these chromatin harbors upon ZGA may release transcription factors and allow them to bind to gene promoters and other regulatory elements to facilitate genome activation. Taken together, these data not only reveal a conserved mechanism underlying chromatin transition during mammalian ZGA but also advanced our understanding about epigenome reprogramming during human in vitro fertilization and early development.
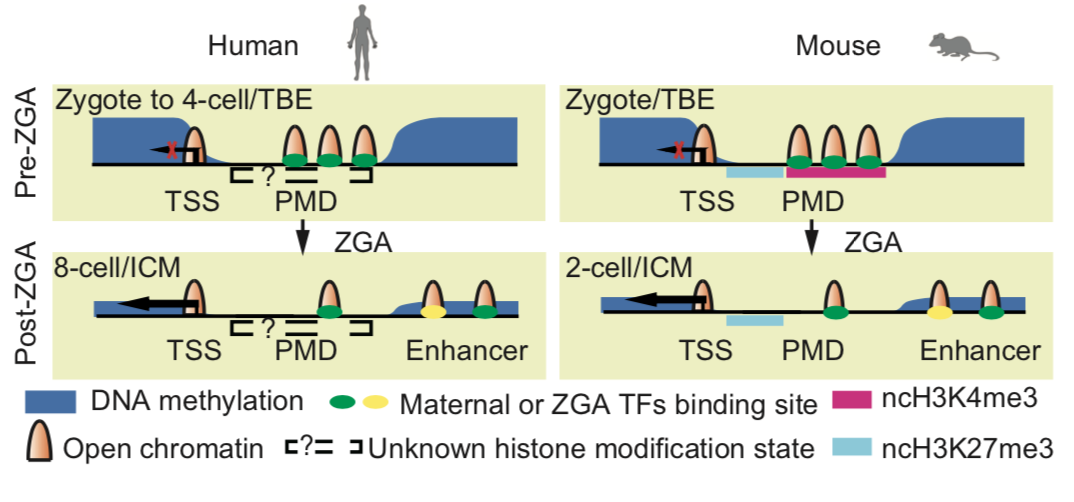
A model shows the distinct transcription and chromatin states before and after ZGA in human and mouse
Prof. Yingpu Sun from Center for Reproductive Medicine the First Affiliated Hospital of Zhengzhou University, PI Wei Xie from CLS, and Prof. Jie Na from School of Medicine at Tsinghua University are the co-corresponding authors of this work. PhD student Jingyi Wu, from PTN program of School of Life Sciences at Tsinghua University, PhD student Bofeng Liu, from CLS program of School of Life Sciences at Tsinghua University, postdoc fellow Zili Lin, PhD student Peizhe Wang, from School of Medicine at Tsinghua University, Dr. Jiawei Xu and Dr. Guidong Yao from Center for Reproductive Medicine of the First Affiliated Hospital of Zhengzhou University are the co-first authors of this work. Collaborators include Professor Wei Li and graduate student Xuepeng Wang from Institute of Zoology, Chinese Academy of Sciences. Bo Huang from Peking University also made important contribution to the study. The study was supported by the funding from the National Key R&D Program of China, National Basic Research Program of China (973 program), the National Natural Science Foundation of China, the funding from the Center for Life Sciences and the HHMI International Research Scholar Award, and also by the animal facility, the sequencing facility and the computation facility at Tsinghua University.
Paper link: https://www.nature.com/articles/s41586-018-0080-8



